General biochemical background. The genetic information on how to build proteins able to perform different biological/biochemical functions is encoded in the DNA sequence. Code-words of three letters/bases, called triplets or codons, are used to encode the information that will be used to synthesize proteins. Every codon encodes the information for one amino acid and every amino acid can be encoded by one or more codons. The genetic code is the biochemical system that establishes the rules by which the nucleotide sequence of a protein-encoding gene is transcribed into mRNA codon sequences and then translated into the amino acid sequences of the corresponding proteins. The genetic code is an extension of the four-letter alphabet found in DNA molecules. These “letters” are the DNA bases: Adenine, Guanine, Cytosine, and Thymine, usually denoted A, G, C, and T respectively (in an RNA molecule, T is changed to U, uracil).
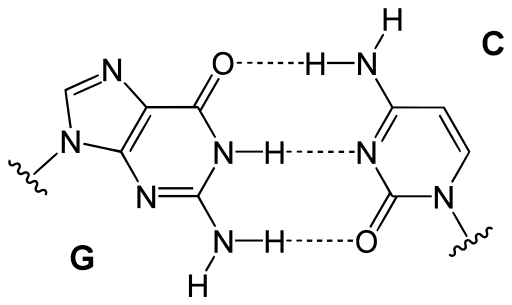
In the DNA molecules, nucleotide bases are paired according to the following rule (Watson-Crick base pairings): G:C, A:T. That is, base G is the complementary base of C, and A is the complementary base of T (or U) in the DNA (or in the RNA) molecule and vice-versa.
Each DNA/RNA base can be classified into three main classes according to three criteria: chemical type (purines: A and G, or pyrimidines: C and T), number of hydrogen bonds (strong or weak) and the whether the base has an amino group or a keto group [1]. Each criterion produces a partition of the set of bases [2].
The relationships between the DNA nucleotide bases, quantitatively expressed throughout the Watson-Crick base-pairings, permit the representation of the standard genetic code as a cube inserted in the Euclidean three-dimensional vector space $\mathbb{R}^3$ [3,4]. In particular, it is plausible that the present standard genetic code was derived from an ancestral code architecture with five or more bases (see main text for full discussion). The algebraic and biological model suggests the plausibility of the transition from a primeval code with an extended DNA alphabet $\mathfrak{B}$ ={D,A,C,G,U} to the present standard code, where the symbol “D” represents one or more hypothetical bases with unspecific pairings. It is important to observe that though the evidence from organic chemistry experiments supports the necessity of five or more DNA bases in the primordial genetic system apparatus, the formal development of the algebraic theory necessarily leads to an extension of the DNA base alphabet. In fact, the additional base was implicit in the multiple sequence alignments of the DNA sequences as gaps representing insertion and deletion mutations (indel mutations). The importance of considering the indel mutations in the phylogenetic analysis was analyzed in [2]. Perhaps the most significant role of the fifth base in the current DNA molecules is played by the epigenetics role of cytosine DNA methylation (CDM). CDM patterning represents one feature of the epigenome that is highly responsive to environmental stress and associates with trans-generational adaptation in plants and in animals.
If the Watson-Crick base pairings are symbolically expressed by means of the sum “+” operation in such a way that the following relationships hold: G + C = C + G = D, and A + U = U + A = D, then this requirement leads to the definition of an additive group or Abelian group on the set of five RNA (DNA) bases. Explicitly, it will be required that bases with the same number of hydrogen bonds in the DNA molecule and different chemical types be algebraic inverses of each other in the additive group defined on the set of DNA bases. In other words, the complementary RNA bases G:C and A:U (or G:C and A:T in the DNA) are, respectively, algebraic complements. This definition also reflects the non-specific pairings of the ancient hypothetical base(s) D, which is taken as the neutral element of the sum operation. Next, there is only one possible definition for the multiplication operation ($\times$) (with base A as the neutral element for this operation) in such a way that it completes a finite (Galois) field structure isomorphic to the field $\mathbb{Z_5}$ and defined over the set of integers modulo 5 (GF(5)). The simplicity of these operations can be noticed in Table 1.
Table 1. Operation tables of the Galois field (GF(5)) on the ordered set of the extended bases alphabet $\mathfrak{B}$={D, A, C, G, U}, and on $\mathbb{Z_5}$.
| Sum | Product | |||||||||||
| + | D | A | C | G | U | × | D | A | C | G | U | |
| D | D | A | C | G | U | D | D | D | D | D | D | |
| A | A | C | G | U | D | A | D | A | C | G | U | |
| C | C | G | U | D | A | C | D | C | U | A | G | |
| G | G | U | D | A | C | G | D | G | A | U | C | |
| U | U | D | A | C | G | U | D | U | G | C | A | |
| Sum | Product | |||||||||||
| + | 0 | 1 | 2 | 3 | 4 | × | 0 | 1 | 2 | 3 | 4 | |
| 0 | 0 | 1 | 2 | 3 | 4 | 0 | 0 | 0 | 0 | 0 | 0 | |
| 1 | 1 | 2 | 3 | 4 | 0 | 1 | 0 | 1 | 2 | 3 | 4 | |
| 2 | 2 | 3 | 4 | 0 | 1 | 2 | 0 | 2 | 4 | 1 | 3 | |
| 3 | 3 | 4 | 0 | 1 | 2 | 3 | 0 | 3 | 1 | 4 | 2 | |
| 4 | 4 | 0 | 1 | 2 | 3 | 4 | 0 | 4 | 3 | 2 | 1 | |
Further readings are provided in the Computable Document Format (CDF) named IntroductionToZ5GeneticCodeVectorSpace.cdf. In this CDF, readers will gain a better comprehension of the biological and algebraic background on the genetic code algebras presented in reference [4]. This is a didactic and interactive visualization to introduce the Genetic code $\mathbb{Z}_5$-Vector Space.
- Cornish-Bowden, A. (1985). Nomenclature for incompletely specified bases in nucleic acid sequences: recommendations 1984. Nucleic Acids Research, 13(9), 3021–3030.
- Jimenez-Montano MA, de la Mora-Basanez CR, Poschel T. The hypercube structure of the genetic code explains conservative and non-conservative aminoacid substitutions in vivo and in vitro. Biosystems, 1996, 39:117–25
- Sanchez R, Grau R, Morgado E (2006) A novel Lie algebra of the genetic code over the Galois field of four DNA bases. Math Biosci 202: 156-174.
- Sánchez R, Grau R (2009) An algebraic hypothesis about the primeval genetic code architecture. Math Biosci 221 : 60 – 76.